Machine Learning in Genomics & Cancer Treatment
When I first arrived in the RxDS (RxDataScience) Headquarters here in the Research Triangle Park as a Machine Learning researcher, I was informed that my first task would be to ‘Redefine Cancer Treatment’. As I’m sure you can understand, coming from a scientific background my curiosity levels were sky high. I was told that the key to bringing personalized medicine to cancer could be found within the genomes of cancer patients. Therein lies a multitude of mutations and variants, with some being benign passengers in this journey and others the core malignant drivers of the cancer itself (McFarland, Mirny and Korolev, 2014). Successfully interpreting these mutations and variants, using either traditional methods or contemporary data science solutions could lead to new treatments with each giving patients with the same underlying causes a fighting chance to overcome their individual cancers.
Machine Learning is a relatively new approach to solving a centuries-old problem. But with the rapid growth of Machine Learning tools, such as Tensorflow, H2O and others coupled with high-end hardware such as NIVIDIA GPU Cards (using CUDA) running on AWS (Amazon Web Services Cloud), it is possible that we may be able to make yet unknown discoveries using the full potential of software and hardware combined!
The Intricacies of Cancer
Developing new therapies to eradicate cancer has never been an easy task, particularly due to the fact there is an array of distinct diseases which all fall under the umbrella term of ‘cancer’, all of which have their own methods of causing a cell’s day-to-day functions go awry (National Cancer Institute, 2018). Having said that, all these diseases do share the same core hallmarks – from self-sufficient growth signals and insensitivity toward anti-growth signals to escaping death (apoptosis) and immortality (unlimited replicative capacities) (Hanahan and Weinberg, 2011). After reading this I’m sure you’re wondering how genetics could play a role in something as destructive as this.
Genetics gone wrong
To start, genetics can be simply defined as the study of heredity (Hartl, 2014). The genes that carry this hereditary information are the building blocks of life and should they become misinterpreted or compromised, chaos ensues. This is where mutations can create the missing link between genetics and cancer. When your DNA is damaged, it becomes mutated. When a gene becomes mutated, it outputs abnormal information to the cell via its resultant protein. This abnormal protein may encourage the cell to divide relentlessly, thus leading to a cancerous state (National Cancer Institute, 2018).
Linking Genetics and Cancer
Now as we’ve all been told many a time and oft, both exposure to the sun’s ultraviolet rays and smoking are primary causes of cancer. What you may not have known is that they’re such potent carcinogens due to how damaging they are to DNA (Bertram, 2000). This is how mutations are derived which have the potential to initiate a cancer in one’s body.
Gaining Ground on Cancer
It’s not all doom and gloom however. Thanks to pioneering research in the fields of technology and the pharma/genomics industries therein, we have staggering amounts of genetic information at our disposal and the tools to interpret it (Behjati and Tarpey, 2013). Behind every newfound genetic mutation or variant linked to cancer thus far, lies a man or woman researching the disease and publishing their invaluable notes in research papers available online – and while we may have catalogued a myriad of genetic mutations as potential drivers, there are still an untold number of variants still out there (Vogelstein et al., 2013). Furthermore, should a skilled oncologist request a genetic sample of a patient’s tumourous cell and then go on to highlight thousands of potential drivers within the sample, he or she must then pore over numerous documents trying to analyse any evidence related to each of these variants in order to accurately classify them. This painstakingly slow task is where I intend on stepping in.
The Task at Hand
The genetic mutations available to us today have all been annotated and categorized into nine distinct classes based on their oncogenicity (i.e. how likely they are to promote cancer growth). The standards have been set by the American College of Medical Genetics and this has all been compiled for us in the Kaggle competition “Personalized Medicine: Redefining Cancer Treatment” which was hosted by MSKCC in an attempt to create competition between data scientists across the world to develop the best machine learning algorithms to discover the class of genetic variants by training up their algorithms on real world evidence (RWE) clinical data. Readily available healthcare datasets from sites such as CMS can also be used in conjunction with such data for feature engineering to enrich the dataset. The field of cancer research, or oncology, is arguably one of the best areas of applying machine learning in healthcare, using neural networks and other predictive analytics algorithms that can parse complex data!
Machine Learning – Breaking it Down
Defining the very essence of what an algorithm truly is makes for no easy task and is still a hot topic amongst some of the brightest minds out there today (Blass and Gurevich, 2003). However, for the uninitiated, an algorithm in its most fundamental form is a step-by-step set of instructions designed for a sequence of operations (Stone, 1972). Machine learning (or ML; an application of artificial intelligence) is built upon the idea that these very same algorithms should be able to access data and use it learn for themselves (Samuel, 1959). The uprising of Machine Learning can be seen in our day-to-day lives and the thought of bringing ML to the burgeoning field of genetics is not an unheralded one, as current efforts are already at the frontier of many disciplines such as genome sequencing and gene editing (Libbrecht and Noble, 2015) (Hough et al., 2016).
Intertwining Genetics and Machine Learning
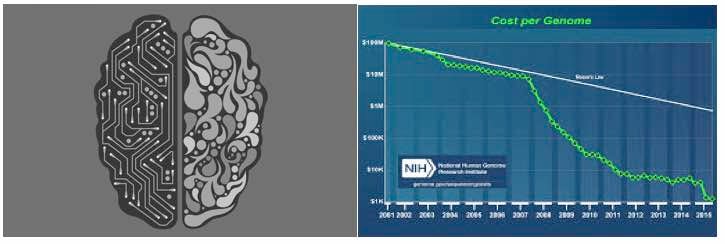
It’s no secret that when it comes to ML, the more complex a question you’re asking, the more a bigger dataset can benefit you (Junqué de Fortuny, Martens and Provost, 2013). For this reason alone, genetics was never a strong contender for potential ML studies due to the scarcity of genetic data available to us. That all changed for the better, however, with the dawn of Next Generation Sequencing (NGS). NGS is the catch-all term for the most modern efforts to sequence DNA at unparalleled speeds - efforts which have been wildly successful (Behjati and Tarpey, 2013). This revolution in high-throughput sequencing has brought the prices of sequencing to all-time lows, making genetic sequencing more accessible than ever and thus opening the floodgates for applications of genomic data in ML.
Final Words
All of the above has culminated to create the foundations of my task here at RxDS. Following on from this, next week I’ll dive into the technical aspect of the task at hand, learning more about the Random Forest algorithm and how we can utilize this with the datasets we have to search for connections between the real world evidence of both analyzed and non-analyzed clinical reports to discover more about the story a patient’s genetics is telling us about their cancer.
About the Author
Conor Moran is a graduate of Genetics from University College Dublin and an aspiring data scientist. He is exploring the capabilities of machine learning within the life science disciplines where he is hoping to make a significant impact in years to come. Conor is currently working alongside RxDataScience to spearhead the efforts of changing the way we think about our genetic data from a clinical perspective.
References
- McFarland, C., Mirny, L. and Korolev, K. (2014). Tug-of-war between driver and passenger mutations in cancer and other adaptive processes. Proceedings of the National Academy of Sciences, 111(42), pp.15138-15143.
- National Cancer Institute. (2018). What Is Cancer?. [online] [Accessed 8 Mar. 2018].
- Hanahan, D. and Weinberg, R. (2011). Hallmarks of Cancer: The Next Generation. Cell, 144(5), pp.646-674.
- Hartl, D. (2014). Essential genetics. Burlington, MA: Jones & Bartlett Learning.
- Bertram, J. (2000). The molecular biology of cancer. Molecular Aspects of Medicine, 21(6), pp.167-223.
- National Cancer Institute. (2018). [online] Genetics. [Accessed 8 Mar. 2018].
- Behjati, S. and Tarpey, P. (2013). What is next generation sequencing?. Archives of disease in childhood - Education & practice edition, 98(6), pp.236-238.
- Vogelstein, B., Papadopoulos, N., Velculescu, V., Zhou, S., Diaz, L. and Kinzler, K. (2013). Cancer Genome Landscapes. Science, 339(6127), pp.1546-1558.
- Kaggle. (2018). Personalized Medicine: Redefining Cancer Treatment | Kaggle. [online][Accessed 8 Mar. 2018].
- Blass and Y. Gurevich. Algorithms: A quest for absolute definitions. Bulletin of the European Association for Theoretical Computer Science, 81:195–225, 2003.
- Stone, H. (1972). Introduction to computer organization and data structures. New York: McGraw-Hill, [1971, c1972].
- Samuel, A. (1959). Some Studies in Machine Learning Using the Game of Checkers. IBM Journal of Research and Development, 3(3), pp.210-229.
- Libbrecht, M. and Noble, W. (2015). Machine learning applications in genetics and genomics. Nature Reviews Genetics, 16(6), pp.321-332.
- Hough, S., Ajetunmobi, A., Brody, L., Humphryes-Kirilov, N. and Perello, E. (2016). Desktop Genetics. Personalized Medicine, 13(6), pp.517-521.
- Junqué de Fortuny, E., Martens, D. and Provost, F. (2013). Predictive Modeling With Big Data: Is Bigger Really Better?. Big Data, 1(4), pp.215-226.


